News and Trends 1 Feb 2023
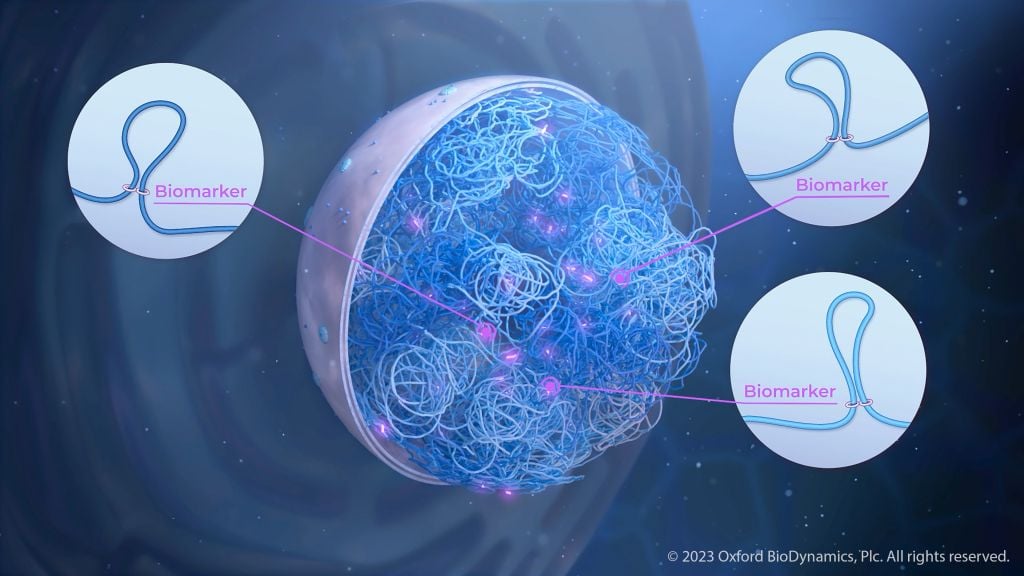
Advancing cancer care with 3D genomics
By Oxford BioDynamics Cancer care is in urgent need of effective tests to determine the best approach for personalized medicine. Practical, new solutions exploiting the power of 3D genomics are emerging. Cancer is a vastly complex, multidimensional disease marked by the uncontrolled proliferation of cells. Many intertwined layers of biological systems give rise to the […]