There’s a growing appetite to refine and standardize CRISPR protocols, as its application expands into different areas of genomic research. The Wellcome Trust and Desktop Genomics have recent advances to make it more efficient and controllable.
Outside of the high-stakes world of therapeutic CRISPR, which has enjoyed gambling-like investment and whose legal future is unfolding in court, this revolutionary technology has also become a hit in genomic research that explores the roles of genes in development and diseases. To aid such studies, academia and industry are developing CRISPR platforms that can make gene editing more reliable.
One example is the University of Cambridge and the Wellcome Trust Sanger Institute. Now published in Development, their teams developed two freely available methods for CRISPR systems, which can be used together to better study genes’ functions.
sOPiTKO is a knockout system that turns off genes by disrupting the DNA, while sOPTiKD is a knockdown system that silences the action of genes by disrupting the RNA. A major advantage is that this can be applied at different stages of tissue differentiation, instead of just seeing the consequences of a gene’s absence at the very beginning of cell development. This opens new opportunities to study the role of genes in time.
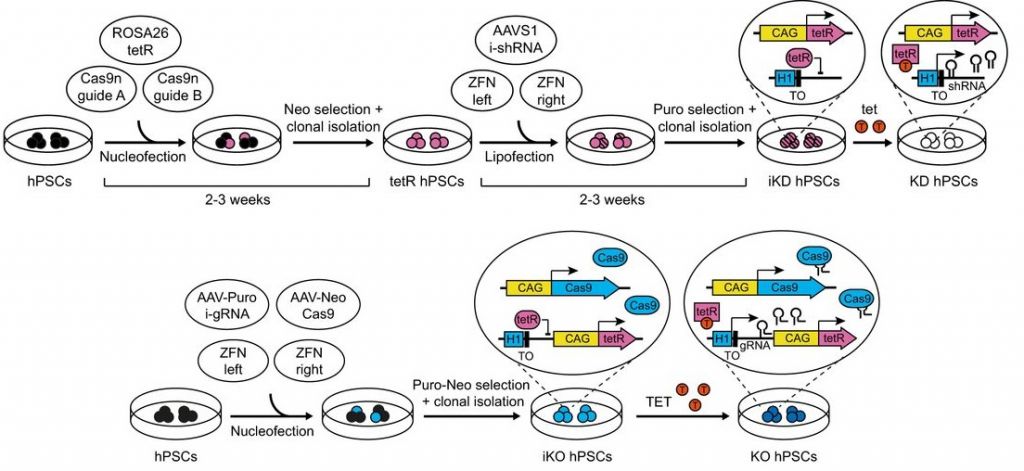
As a single-step system with the ability to turn off entire families of genes, sOPiTKO and sOPiTKD could have a big impact in the fields of developmental biology, tissue regeneration, and cancer. Their developers estimate that its improved efficiency and reliability could cut a 9-month experiment down to just a few months for a typical study of the activity of 3 genes.
However, this is far from being the only effort in standardized CRISPR platforms. Last week, a partnership between Desktop Genetics and LGC, a multinational in the life sciences space, was awarded a grant by SynbiCITE, a UK center for the commercialization of synthetic biology. The goal is to create standard protocols to use CRISPR, namely to detect and quantify the results of the targeted mutations. Having such a tool would greatly help validate the work of labs using CRISPR, and thus usher in more discoveries like this mysterious immunology mechanism.
Images: Khakimullin Aleksandr (Shutterstock ID535724044); Bertero et al. (2016) Optimized inducible shRNA and CRISPR/Cas9 platforms for in vitro studies of human development using hPSCs. Development (doi: 10.1242/dev.138081)





