This is the story of how the Natural History Museum of Denmark and Molecular Ecology biotech Spygen (France) brought together forensic science techniques to detect traces of Great Crested Newt DNA in UK ponds.
The Great Crested Newt (Triturus cristatus) is an amphibian found across Northern Europe and parts of Asia, which has been declining since the 1940’s due to loss of habitat and is a well-known UK protected species.
But how can we monitor how many of each species are in a particular habitat – and whether certain areas need additional legislative protection?
Previously, the detection of species was done through exhaustive hunts for eggs using bottle-trap set-ups (over several months), and night-time torchlight surveys, which are challenging (and have a lot of room for inaccuracy)!
Therefore, funded by the UK Department for Environmental and Rural Affairs (DEFRA), Natural England, the Joint Nature Conservation Committee and Scottish Natural Heritage, a novel method of detecting the species was developed in 2013 using eDNA.

Described as a tool for ‘Biodiversity CSI‘, environmental DNA (eDNA) is genetic material released into the water by animals from their skin, feces, mucous, hair, eggs and sperm. Measuring species presence with eDNA therefore only needs a single pond water sample, then used to test for traces of newt DNA.
Spygen is a French biotech specializing in molecular ecology, and a 2011 spin-off from the Laboratoire d’Ecologie Alpine (LECA), the CNRS at the Université Grenoble and the Université Savoie Mont Blanc.
The UK project was therefore undertaken by the Freshwater Habitats Trust using Spygen’s eDNA tools, also collaborating with Amphibian and Reptile Conservation and the Durrell Institute of Conservation and Ecology at the University of Kent.
Samples of eDNA were analysed using primers and probes designed by a team from the centre of GeoGenetics at the Natural History Museum of Denmark in Copenhagen. They also wrote a great review on the potential of eDNA for monitoring biodiversity.
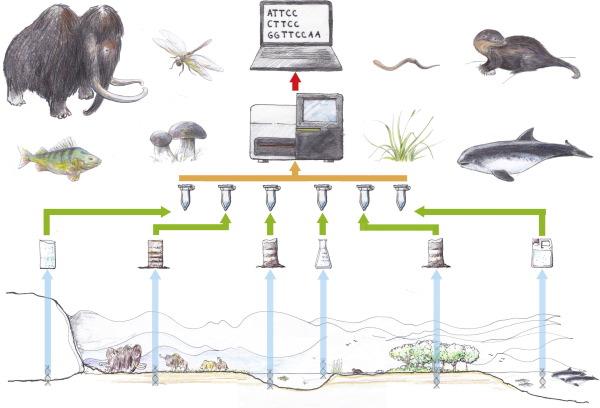
During the research, in silico analysis was first performed with ecoPCR software, comparing primer-template analysis from the EMBL-Bank and the SPYGEN reference database which includes 56 European amphibian species.
Primers and probes were tested in vitro against tissue samples collected from 16 individual great crested newts by external swabbing from three different populations. DNA was finally extracted using the DNA Blood and Tissue kit developed by the European NGS specialist, Qiagen (Germany).
The eDNA method was found to be highly effective at detecting the presence of Great crested newts – even more effective than traditional sampling methods. It is also proving to be an easier, non-invasive and cheaper alternative (all significant advantages in conservation).
This Molecular biodiversity tool therefore has benefits for the Newts, as well as businesses and land-owners who need to monitor the species presence on their property. Once again, thanks Biotech!
Feature Image Credit: Remix of Graphics by Labiotech (Source: Spygen)
Figure 1:Thomsen, P. & Willerslev, E. (2015) ‘Environmental DNA – An emerging tool in conservation for monitoring past and present biodiversity’, Biological Conservation, 183(4-18). doi:10.1016/j.biocon.2014.11.019





