Newsletter Signup - Under Article / In Page
"*" indicates required fields
A team of French scientists has built the first molecular computer using polymers to store data, making each bit 100 times smaller than with current data storage.
Just eight bytes encoding the word “Sequence” in ASCII code. A feat for the team of researchers that encoded and read the word using a synthetic polymer sequence. Their experiments, published today in Nature Communications, prove that it’s possible to store information in polymer molecules. Given the size of each monomer unit of the molecule, this method makes the storage of each bit of information — either a 1 or a 0 — a hundred times smaller than that of current hard drives.
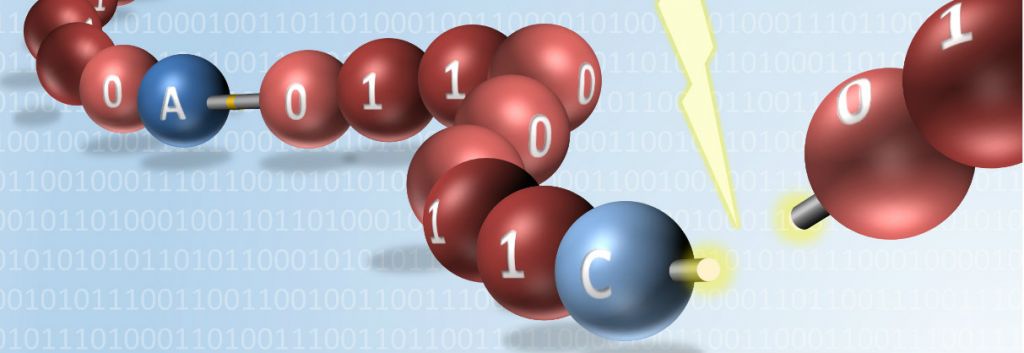
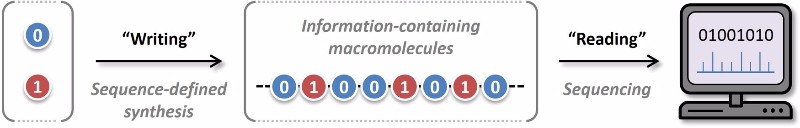
The researchers, from the Institut Charles Sadron (CNRS) in Strasbourg and the Institute of Radical Chemistry (CNRS) in Marseille, devised a method to encode information into polymers by using two different types of monomers containing phosphate groups. One type corresponded to 1 and the other to 0. After every eight monomers, or “bits,” a molecular separator was introduced to separate each byte.
In order to read the information encoded in the polymer, each byte is cut at the separator site. Then, the phosphate bonds between the monomers are broken, and the eight monomers are identified using a mass spectrometer.

Although it took hours to encode and read the information, the researchers are confident that the method can be dramatically sped up by automating polymer synthesis and sequence analysis. Their next goal is to build the first molecular floppy disk by making a larger molecule that can store several kilobytes of data, roughly the equivalent of a page of text.
With the huge speed at which data storage is growing, many are searching for new methods to replace silicon computers and store data more efficiently and using less space. Quantum computing and DNA computing have been gaining a lot of traction, but it currently not possible to build these computers at a large enough scale to substitute silicon. Microsoft itself is working on improving DNA computers, while the EU is investing in researching other biocomputing alternatives based on motor proteins.
The creators of the polymer-based molecular computer argue that a key advantage of their method is that synthetic molecules, with structures that can be optimized for mass spectrometry, are much simpler to work with natural biomolecules such as DNA. Still, I’m sure they will have a lot of challenges to overcome in order to scale-up, for example, the fact that mass spectrometry requires destroying the molecules analyzed.
Images via Jean-François Lutz /Institut Charles Sadron; Macromolecules, 2015, 48 (14), pp 4759–4767