Newsletter Signup - Under Article / In Page
"*" indicates required fields
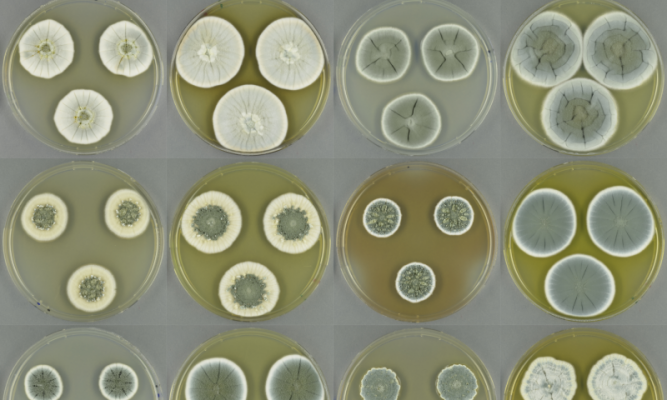
Good old Penicillium still has a lot more tricks upon its sleeve: Swedish researchers see its potential as a gold mine for new antibiotics.
The genomic analysis of Penicillium species represents a potential gold mine for new bioactive compounds, or improved versions of those we already use. Published recently in Nature Microbiology, the work was carried at Chalmers University of Technology, Sweden, where there is an overarching effort to study fungi’s potential as new cell factories. The effort has led to a new method of antibiotic discovery, a useful tool in the fight against antibiotic resistance.
Fungi, and more specifically Penicillium strains, have long been very useful in biotech. For the area of health, they are used for the industrial production of the antibiotic penicillin as well as statins, cholesterol-lowering drugs. With such demonstrated potential in mind, Chalmers’ researchers thought it would be fruitful to sequence the genome of nine Penicillium species.
Not unlike the NGS of human genomes, bioinformatic analysis and interpretation of all the data produced by sequencing are key to making it useful. These researchers are using a method to identify the genes coding for compounds of interest, particularly those that are yet unknown.
How does one look for something that isn’t yet known? Most of these interesting bioactive compounds are secondary metabolites, which include antibiotics and many flavoring compounds. Rather conveniently, the genes coding for such compounds tend to cluster together and have some signature characteristics. This leads researchers to pathways yet without a known function, but with a higher probability to code for interesting molecules.
The first success of this sequencing and analysis was the discovery of a new kind of yanuthone, a class of molecules that have shown anti-fungal activity. The results also turned up more producers of a known yanuthone, and this type of findings can be helpful to find better strains to produce known compounds.
This kind of “mining” has been done before in many bacteria, including extremophiles of remote areas, but the authors of the study say fungi remained to be tackled. The study shows a vast range of secondary metabolite pathways, and a lot of them are yet to be explored.
The urgency of exploring such pathways is due in big part to the need of discovering new antibiotics. Antimicrobial resistance is on the rise, with some very grim forecasts, and the antibiotic research has been neglected by pharma in the past decades. While there has been a revival of interest in biotech, there is still a lot to be done. Mining the genomic biodiversity of more complex organisms, with quicker methods, appears to be an important way forward.
Images by MDWines/Shutterstock and Jens Christian Nielsen
Partnering 2030: FME Industries Report







